We were excited to see Ben Davis’ group publish their first paper with their new Franklin address earlier this month. Working with Emilio Cocinero from Universidad del Pais Vasco (UPV) and Francisco Corzana from Universidad de la Rioja, Ben Davis’ group have successfully used a new method to determine the configuration of “naked” sugar molecules, known as deoxyribosugars which form part of DNA’s building blocks.
Understanding of these isolated, ‘naked’ structures is crucial, as their composition, shape and flexibility of these structures can control their biological function. There is growing interest in the design of these DNA building blocks, as both naturally occurring and man-made structures can have diagnostic and therapeutic applications.
To observe the structure of the ribosugars in an unaltered state, they had to be generated and analysed using novel methods. This novel method utilised new spectroscopic instrumentation developed by Professor Cocinero; details of the method can be found in their recently published work in ACS Central Science journal.
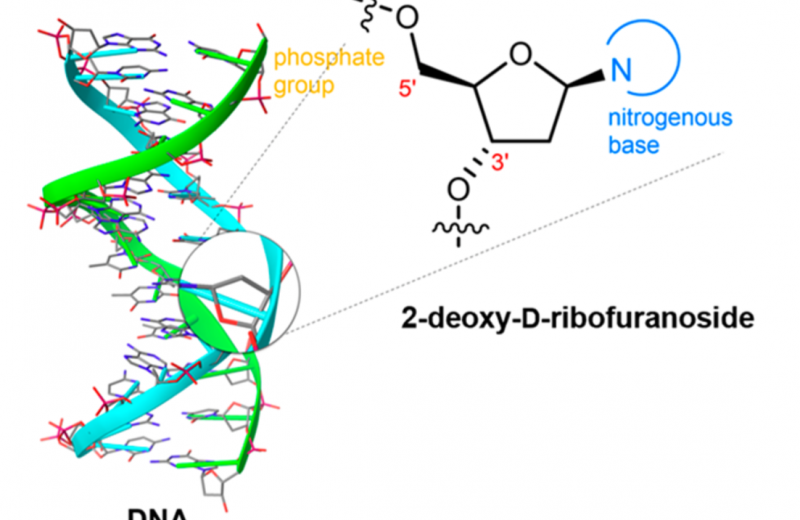
Gas Phase Spectroscopy
This spectroscopic method allowed the determination of the true, high resolution structures of deoxyribosugars. This is possible, in part, due to the detection of slight variations in number of neutrons present in the atoms which make up the sugar molecules. These molecules with naturally differing number of neutrons are known as isotopes, for example, carbon has three isotopes carbon-12, carbon-13 and carbon-14. The differing number of neutrons in these isotopes does not alter the chemical behaviour of the carbon atoms, but means they can be differentiated due their different molecular weights because the heavier atoms move slower than the lighter atoms.
The team were able to synthetically construct different deoxyribosugar scaffolds and to accurately detect these small differences in the mass of these molecules because their custom-made design of microwave apparatus used in the spectrometer, which allowed for a superior frequency and chemical resolution. An additional advantage of this method, is that the increased sensitivity and prolonged time over which the sample is tested, means that a smaller sample is required.
Prior to this work, such high-resolution structures of deoxyribosugars could not be observed in an isolated, unaltered state. Our understanding of the DNA structure mainly comes from X-ray crystallography, which requires the DNA to be a crystalline solid before determining the structure.
While this process has been invaluable for determining the helical structure, base-pair stacking and some backbone conformation, it only provides information about certain static structures and not different possible shapes. For this, the dexoyribosugars may be better observed in a gas phase so that different conformations can be caught.
Prior advances in spectroscopy have previously allowed for analysis of some arrangements of sugars in their gas phase. However, during this method, a tag often needs to be added to the molecules to aid with their detection, but this addition can induce changes in the structure and cause unwanted environmental interferences. This latest advancement in technology has now allowed Ben’s team to view the configuration of deoxyribosugars in their gas phase without the need to attach these tags and to view the ‘naked’ shapes.
DNA Sugar Structures
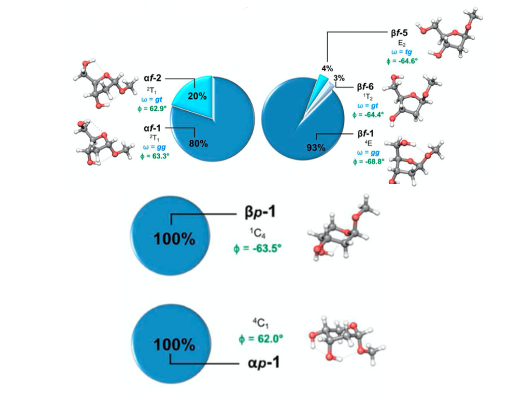
The new method allowed for an extensive structural ‘shape’ map of several possible ribosugar scaffolds to be built. The conformations of two forms of the furanosides (five-membered rings, containing four carbon atoms and an oxygen) and two forms of the pyranosides (six-membered rings, containing five carbon atoms and an oxygen) were determined, pictured below with their population and key geometrical parameters. For the furanosides five conformations were found, two for the α form and three for the β. While, for the pyranosides both forms (α and β) had a single dominant conformations.
Understanding the relative prevalence of each of these shapes can help us elucidate the preferences of these nucleotides, three of which are unnatural and one natural. The natural scaffold appears to display shape preferences, now observed in this paper, that may give a clue to why this has been predominantly utilized as the sugar backbone for DNA. This deeper insight into these preferences can in turn aid in our understanding of natural nucleotide function and the development of man-made, unnatural nucleotides.
Related publication
Calabrese et al. - Observation of the unbiased conformers of putative DNA-scaffold ribosugars - ACS Cent. Sci., 6, 2, (2020)