Mapping the Developing Hippocampus with Synapse Safari
Scientists at the Franklin have teamed up with the MRC Centre for Neurodevelopmental Disorders at King’s College London to deliver our next citizen science project, Science Scribbler: Synapse Safari.
Grab your virtual microscope and get ready for an adventure with Science Scribbler: Synapse Safari! Help us outsmart AI as we venture into uncharted depths of the developing hippocampus.
This project aims to take a closer look at the synapses, the sites where neurons connect and transmit information, and study the mitochondria and synaptic vesicles they contain. Volunteers are asked to identify and outline these structures in a process called segmentation. But for the first time in Science Scribbler history, instead of doing so from scratch, volunteers will be verifying and correcting the segmentation results obtained from machine learning – a process known as proofreading.
Researchers at KCL acquired images of mouse hippocampus using serial block-face scanning electron microscopy, and manually annotated the data volumes to segment the synapses. Then, Franklin scientists applied two machine learning approaches to segment the mitochondria and vesicles within the expert-segmented synapses.
Synaptic vesicles are relatively uniform in size and shape, making them suitable for a technique called template matching. In this technique, a small template image of a vesicle is compared to different parts of the larger image to find matches. However, the small size of synaptic vesicles makes them susceptible to noise and imaging artifacts, which can confuse the template matching algorithm.
The mitochondria were segmented using the pre-trained model MitoNet, developed by researchers at the National Institutes of Health [1]. This model has been trained on millions of microscopic images and can accurately segment most mitochondria in the imaging volumes collected for Synapse Safari.
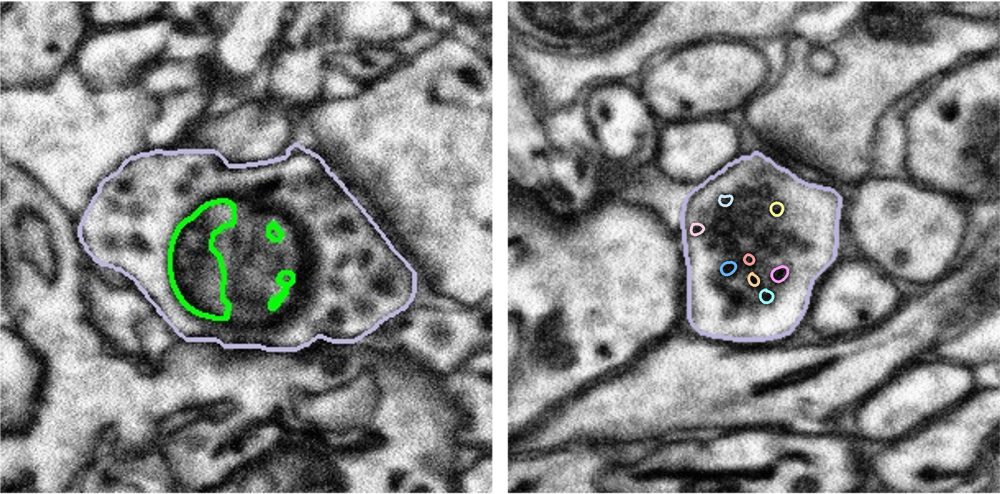
However, even state-of-the-art deep learning models can struggle with the variability and ambiguity of biological images (see image above). Additionally, applying pre-trained models requires inspection and validation of the machine output, and fine-tuning of the model with supplementary segmentation is often needed to improve the model’s performance. As a result, human proofreading and correction remain essential but time-consuming steps in ensuring the quality and reliability of the final result.
The annotations from many citizen scientists will be aggregated together to form a single consensus for each image volume. Ultimately, the researchers hope that knowing the shape, size, number, and location of mitochondria and vesicles within the synapses will contribute to our understanding of how these neuronal connections are functioning in the brain, and might give a hint as to how neurodevelopment disorders like autism and epilepsy can develop.
Find out more and give the project a try on the Zooniverse.
[1] Ryan Conrad and Narayan Kedar, Instance segmentation of mitochondria in electron microscopy images with a generalist deep learning model trained on a diverse dataset, Cell Systems 14.1 2023 58-71
