“We are putting AI at the centre of our biological research, meaning that we are generating the type of data that is needed to drive AI and machine learning. The aim is that, once created, the digital twin cell can be used for virtual experiments, giving researchers a highly valuable and cost effective scientific tool to understand biology and health.
We are training our researchers to develop the skills they need to use AI with ambition and efficiency, thereby expanding AI and machine learning capacity in the UK and accelerating life sciences discovery.”
The ambition
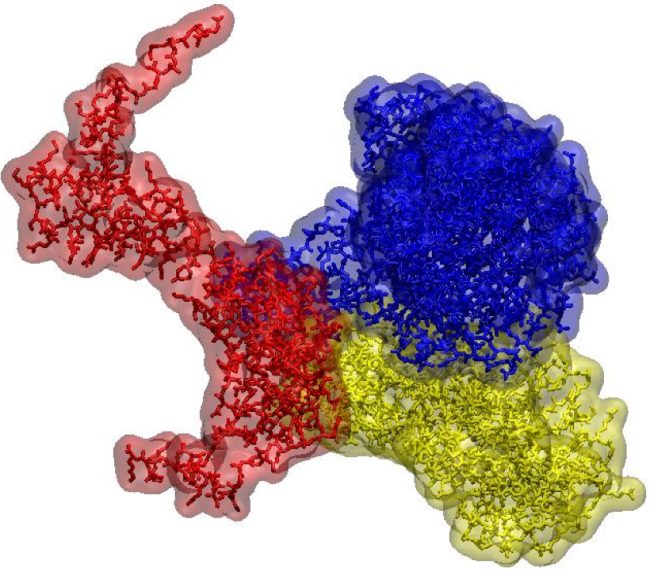
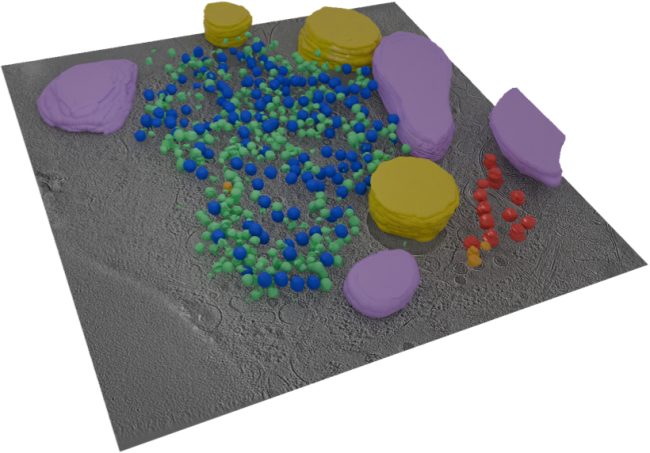
By creating a quantitative digital framework for representing cell ultrastructure and chemistry, we will provide a tool for understanding rules of transformation between cell states and predicting consequences of developmental signalling or pharmacological interactions.
The expertise we build at the Franklin, the open data we share, and the tools we generate, are used by the wider UK research community to boost UK life sciences. The culture we build of sharing high quality data sets with the wider scientific community inspires others to do the same, and drives the generation of revolutionary new tools that change the face of biological research.

What are we doing?
Computer scientists will be an integral part of our matrix research teams tackling complex life sciences challenges. By being involved from the earliest stages of the project, and throughout the whole project lifespan, the highest quality data will be generated and the most appropriate AI and machine learning tools will be used. Being part of these AI best practice project teams, biologists, chemists and physicists themselves will become expert in using AI and machine learning in research.
This Challenge demands exceptionally high quality cryo Electron Tomography (ET) images of cells to be taken in high-throughput on fine lamellae. We will extend methods developed at the Franklin to generate and align digital representations of these samples, making use of the multi-scale data to increase accuracy of reconstruction and segmentation.
Image objects will be annotated in a first pass using tools previously developed for ultrastructure and particles representing proteins and protein assemblies that will be broadly classified by similarity scoring. Refined analyses then will be generated based on additional information (e.g., hierarchical relations, molecular dynamics simulations) for serial cycles of retraining.
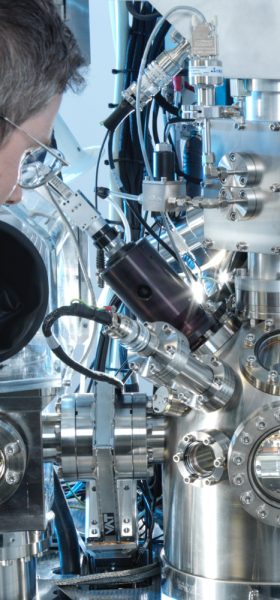
In parallel, we will develop automated methods for preparing the lamellae through “adaptive Focused Ion Beam (FIB) milling” in which the milling is guided by AI from periodically acquired Scanning Electron Microscopy (SEM) images for automated decision-making. These will be applied for acceleration of data collection to generate training sets of the cells in different differentiation states.
Why?
Adapting AI and machine learning for use in biology presents a number of challenges. Biology is complex, shows variations and the data we generate contain margins of error. Harnessing the power of AI and machine learning to understand biology is in its infancy, and as a result is often used inefficiently or in ways that doesn’t yield helpful results.
In addition, too often research projects in the life sciences are designed without input from experts in data, AI and machine learning at an early enough stage. Which can mean that the data generated isn’t optimum for these systems. For example, the data available to AI algorithms may already have been catagorised by researchers, whereas accessing the raw data may have been a better source for machine learning.
Our work in this area will address these challenges and our learning show how AI can be used more efficiently and effectively.
Why the Rosalind Franklin Institute?
Over the past 5 years the Franklin has become internationally recognised for its work around the development of FIB-SEM technologies, specifically pushing the frontiers in plasma FIB work. It has built a suite of tools for CryoET workflows, that are in early stages of allowing full automation of some processes. The Franklin also has pioneered approaches for multi-scale AI-driven segmentation based on crowd-sourced training data and for particle identification.