Researchers identify dozens of new targets for potential COVID-19 drugs
An international, multidisciplinary team of scientists has uncovered a ‘new universe’ of interactions between the SARS-CoV-2 virus and human cells, vastly increasing the potential for new treatments to tackle COVID-19.
Using a method first developed to generate insights into the fever-causing Sindbis virus, the team identified the key cellular proteins affected by SARS-CoV-2’s viral RNA – molecules used by the virus to carry its genetic information into human cells and cause COVID-19 disease.
As a proof of principle, the researchers tested some of these targets with five existing drugs in lab-grown cells infected with SARS-CoV-2. Two of the drugs – cancer therapeutics known as Ganetespib and BTYNB – showed strong effects in suppressing the virus, while two others showed moderate effects.
Professor Shabaz Mohammed of the Rosalind Franklin Institute and Oxford University, one of the study’s senior authors, said the research would ‘massively increase’ the scope for attacking the virus with repurposed drugs, as seen already with Remdesivir.
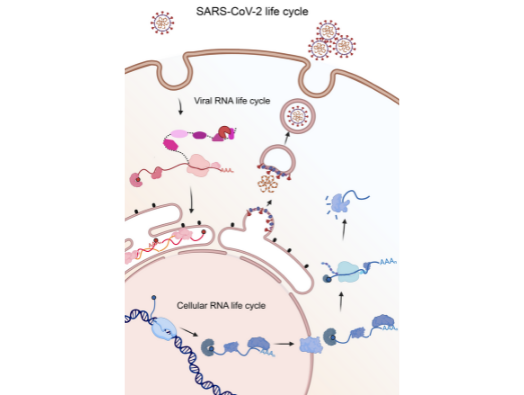
Professor Mohammed, a mass spectrometrist, said: ‘The concept behind this research is that viral RNA essentially hijacks the human cell’s RNA-binding proteins and manipulates them into misbehaving, allowing the virus to spread and thrive. We have been working together as a team for about six years on this idea and saw at the outset of the pandemic that it could be applied to SARS-CoV-2.’
The team’s technique, named viral RNA interactome capture (vRIC), was honed on the Sindbis virus and provides a comprehensive and systematic approach to discovering the RNA-binding proteins that are hijacked by viruses.
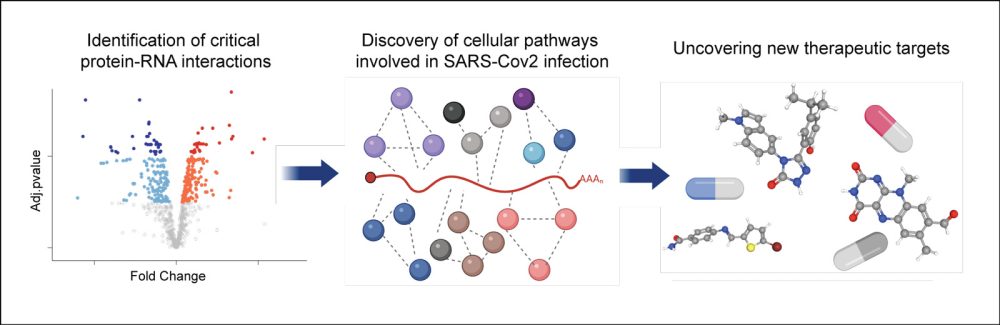
RNA biologist Dr Alfredo Castello, of the University of Glasgow, is another of the study’s senior authors. He said: ‘The Nobel prize-winning biologist Sir Peter Medawar described viruses as “bad news wrapped up in protein”. The bad news is the genome, which in the case of SARS-CoV-2 is made of RNA – that’s the heart of the virus. Through this study we now know which cellular proteins engage with SARS-CoV-2 RNAs, and by cross-referencing these proteins with existing drug databases we found 30 or 40 potential therapeutic targets for COVID-19.
‘Collectively, this is a new universe of host-virus interactions, with hundreds of drugs out there ready to be tried as potential antiviral treatments. Our proof of principle with just five of these drugs found that four had at least some effect in suppressing SARS-CoV-2, which is highly encouraging.’
Crucial to these findings was the vRIC methodology, which relies on advanced high-throughput technology and ultra-sensitive equipment to deliver results quickly and comprehensively. Professor Mohammed said: ‘Developing our methodology over the past six years has made all of this possible, and the technologies being developed at the Rosalind Franklin Institute will have an immense impact on this type of work in the future. It also meant we were immediately able to bring together an international, genuinely multidisciplinary team of researchers to deliver these insights into the coronavirus.
‘Our goal was always to show how these techniques would have real value for translational medicine and virology, and this work on SARS-CoV-2 is a clear demonstration of that. Researchers working on tackling the virus now have a strong list of targets to get their teeth into.’
Dr Castello added: ‘While we will only know the impact of these findings in hindsight, they could be very important for the next phase of the pandemic – for example, in addressing the effects of vaccine-resistant variants or long COVID. Our methodology will also allow us to adapt quickly to any new emerging threats in the future.’
The paper ‘Global analysis of protein-RNA interactions in SARS-CoV-2 infected cells reveals key regulators of infection’ is published in the journal Molecular Cell, read the paper here. The work was led by Dr Wael Kamel and Marko Noerenberg, postdoctoral researchers at Glasgow and Oxford, and Berati Cerikan, postdoctoral fellow at the University of Heidelberg, read more about this research here.
