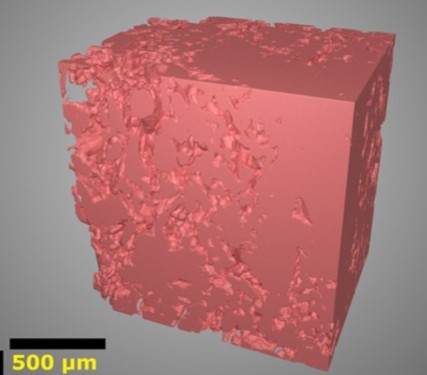
A novel technology for creating 3D images of the placenta’s blood system and understanding its role in high-risk pregnancies
Franklin researchers have developed AI tools able to automatically label different elements of 3D images of human placenta biopsies. With support from a multi-institution collaboration including the Franklin, the researchers created 3D models of the blood vessels within placenta and identified structural features that could limit the transfer of oxygen and nutrients between mother and baby. The team are integrating these models with ultrasound and maternal clinical data to data show how changes in placental structure impact pregnancy conditions including pre-eclampsia, hypertension, maternal diabetes and fetal growth restriction. Based on this experience, the team is creating a tool to help doctors better manage high risk pregnancies to prevent stillbirth.
Background
Worldwide, a baby is stillborn every 16 seconds and, concerningly, stillbirth rates have not changed over the past decade. Stillbirth has multiple causes, however around half of stillbirths result from fetal growth restriction, a pregnancy disorder where impaired placental function severely limits fetal growth. Fetal growth restriction is also a risk factor for other forms of poor pregnancy outcomes such as infection.
The presentation of fetal growth restriction is often a combination of factors arising from the maternal and fetal systems in unique combinations, creating challenges for detection and management. The placenta connects the mother’s and baby’s circulation through separate networks of blood supply which enable transfer of nutrients, oxygen and waste. Due to its key role in facilitating the transfer from mother to baby, the placenta’s structure often reveals the functional changes due to variations in maternal and fetal systems associated with fetal growth restriction.
Current methods of classifying pregnancies as high or low risk are imprecise. Maternal conditions, such as pre-eclampsia, hypertension and maternal diabetes all carry a higher risk of fetal growth restriction and with it stillbirth, however fetal growth restriction is not always detected in utero.
One powerful approach to more accurately distinguish high from low-risk pregnancies is through better understanding of how placental structure and blood flow affect pregnancy outcomes. This can be done through computer modelling – but a lot of data is needed to create a useful model. Generating this data is very time-consuming when done manually, so automated processes using AI tools are needed.
Science in detail
Funded as part of Wellcome Leap’s In Utero programme, which is aimed at reducing stillbirths, Franklin researchers became part of a global team creating digital representations of circulatory systems in the placenta in order to more accurately identify differences in placental structure between high and low risk pregnancies.
Collaborators from the University of Manchester collected patient data, ultrasound and MRI scans from 127 women, From these, it was possible to collect further 3D imaging of the placenta after delivery in 24 cases. Franklin researchers worked with colleagues from the University of Birmingham to carry out synchrotron X-ray computed tomography (sCT) imaging of the placenta samples at Diamond Light Source.
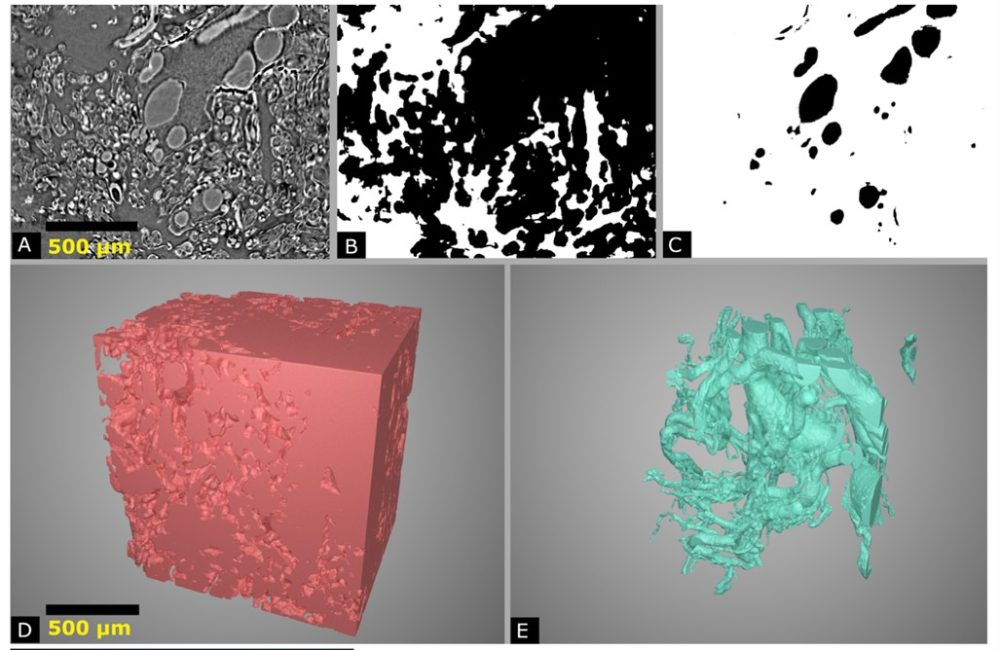

The Franklin team carried out segmentation on these images, which involves labelling the different types of tissue, including fetal vasculature, villous tissue, and the intervillous space, where the maternal blood flows. Using this segmentation, the team generates additional data, including surface area and volume of each tissue type, skeletonisation of the fetal vascular network (e.g. how many branch points and distance between branches), total length and average diameter of the network, and pinch points within the maternal blood space, where pressure could be created or flow might be constricted.
To manage the large amounts of data, the Franklin team created new deep learning approaches [1, 2], to semi-automate this process. Because each placenta is a unique, complex structure, which changes over a pregnancy term, a fully automated deep learning process could not be used. The researchers had to use a bespoke system, initially training the model on each individual 3D volume. They began with small volumes and mainly manual annotation, then scaled up gradually to larger volumes and semi-automated annotation. The models they have developed can now be used on new datasets, with less manual input required. This has enabled the Franklin team to speed up the image segmentation process and move the research forward at pace.
The image data generated by the Franklin team was shared with colleagues in the Universities of Auckland, Manchester, London and Swansea who combined this data with patient data and ultrasound imaging to create models of the placenta, known as ‘digital placentas’, linked to the maternal and fetal circulatory networks to understand how blood flow works across the whole system. This understanding is then validated using the MRI data from each patient, as analysed by colleagues in Cardiff. Together this allowed the team to create a technology that can transform how we interpret ultrasound, to improve the use of this tool in determining pregnancy outcomes.
Why the Franklin?
The researchers were able to draw on earlier work at the Franklin from 2018 which involved segmentation of images from four placenta biopsies samples. The team adapted a sparsely trained but effective computer model developed from this study and added additional training data so it could be used to segment a broader range of datasets.
They were uniquely placed to act as a bridge between the clinical and research teams from partner institutions, given their experience of working in image analysis of biological samples.
The Franklin’s close relationship and proximity to Diamond Light Source helped to facilitate the imaging work on beamline I13-2, drawing on the Franklin’s expertise in image segmentation and creation of deep learning models to segment and manage biological imaging data.
The synchrotron imaging of human placenta collected at Diamond Light Source requires segmentation in order to create useful data. Segmentation is computation hungry and to help with this analysis the Franklin regularly uses Baskerville (University of Birmingham) and also gained early access to the new supercomputer, Isambard-AI (University of Bristol), which sped up the process, enabling science that otherwise would not be possible.
Without the Franklin’s expertise in imaging methods, deep learning models and high-performance computing, the project would not have generated sufficient data to develop a patentable technology.