Crowd-powered research
We have developed open-source tools to support the aggregation, visualisation, and analysis of citizen science data, and shown that citizen science can produce valuable annotations for training and finetuning models for complex 3D biological imaging datasets.
Over 18,000 registered volunteers have contributed 3.49 million classifications to Franklin citizen science projects, totalling 18,000 hours of human effort since 2020. This has accelerated algorithm development, reduced individual effort through aggregation of multiple annotators, and supported public engagement and public participation in Franklin research.
These “Science Scribbler” projects have been showcased at 4 open days, 3 science festivals, and over 50 school workshops. Taking part in these events gives visitors a chance to connect with and contribute to imaging science.

Background
The annotation, segmentation and analysis of large biological volumetric datasets is complex, time intensive, and difficult to automate at scale due to the inherent variability in biological samples. The Franklin has integrated citizen science with AI to diversify and streamline this data pipeline.
The burden of creating manual annotations to use as training data for segmentation models typically falls on the researcher. Not only is this time consuming, but it can lead to bias in the model output, as even expert annotations typically show low levels of agreement because of the noise and complexity of the images.
Citizen science offers an alternative solution that redistributes this work effort to the crowd. By aggregating the annotations from many non-experts, a final segmentation can be produced that is of comparable quality to manual expert annotated methods, but at greater scale and with reduced individual observer biases and the use of less expert annotation.
However, adopting citizen science also poses challenges including developing robust aggregation and cleaning tools for diverse annotation types, validating outputs to build trust in the scientific domain, and building an engaged and committed volunteer community. These all were successfully addressed by the Franklin team.
Science in detail
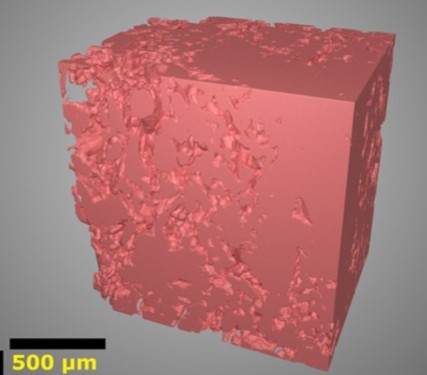
We have implemented Zooinverse Science Scribbler projects using data from SBF-SEM, (S)TEM, cryoSXT and cryoET, targeting the segmentation of organelles (including mitochondria), virus particles, and nanoparticle catalysts. Typical workflows involve point, ellipse or freehand annotations of target structures, followed by a classification and/or cleaning step, and aggregation to generate a single ‘crowd consensus’. Depending on the format, we have found success with aggregation using density-based clustering (e.g. DBSCAN, HDBSCAN), majority voting, and/or height maps with thresholding and outlier removal.
In June 2024 we launched Science Scribbler: Synapse Safari, mapping mitochondria and synaptic vesicles in SBF-SEM images of mouse hippocampus[5]. This project, in collaboration with King’s College London, used a combination of expert and machine segmentation, followed by proofreading of the machine annotations by the crowd.
Mitochondria were initially segmented using MitoNet and synaptic vesicles via template matching. We utilised an experimental Zooniverse feature (“correct-a-cell”) that enables pre-loaded machine annotations to be presented to volunteers, who could delete, amend, or redraw them. The workflow was refined through alpha testing at the Diamond Light Source open day, improving training and task design prior to launch.
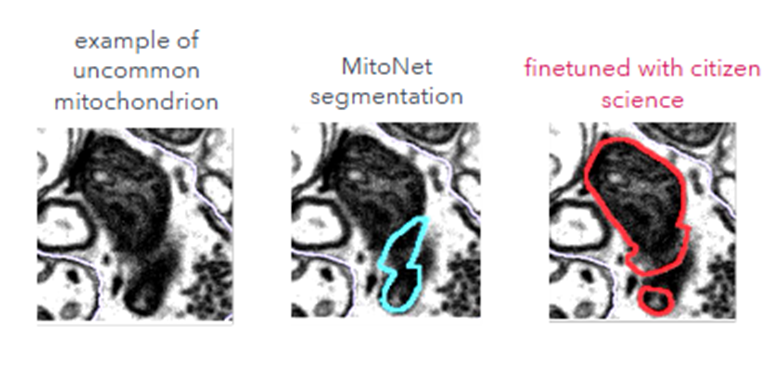
We were able to extend the project beyond its original scope thanks to strong volunteer engagement and segmentation outcomes. In the initial datasets, volunteer corrections of mitochondria inside synaptic boutons were used to finetune MitoNet, leading to a marked improvement in the segmentation of morphologically uncommon mitochondria (F1 score increased from 0.67 to 0.89; IoU from 0.61 to 0.81). The finetuned model was then applied across the full volume. Building on this success, we expanded to SBF-SEM images of 7-day postnatal mouse hippocampus, where volunteers proofread mitochondria sampled across the dataset. These annotations again enabled model finetuning and more accurate large-scale segmentation. In parallel, volunteers generated synaptic vesicle annotations that are now being used to adapt or develop segmentation models suitable for SBF-SEM data with collaborators, where existing synaptic vesicle models have limited applicability.
Impact
Synapse Safari
The volunteer hours taken to complete this comprehensive segmentation is equivalent to one full year of work for an individual researcher making previously unmanageable research tasks achievable.
This comprehensive segmentation of mitochondria enabled by Synapse Safari revealed their previously unreported morphological diversity in the hippocampus. This will be further investigated with molecular imaging techniques, including STORM and SIMS.
In parallel, a large set of high-quality synaptic vesicle segmentations has been generated in the project, which are being shared with collaborators to support the development of improved deep learning tools.
The project also demonstrated the effectiveness of a citizen science workflow based on proofreading rather than de novo annotation. This is a more efficient approach that can be adapted by other biological image analysis projects.
Tools for preparing and processing citizen science data for segmentation
The Franklin has developed several open access tools for the preparing and processing of citizen science data for segmentation. The Franklin team has made it easier for other science teams to enabling other teams to make effective use of citizen science by making a range of tools available, including;
- SuRVoS2 is a napari plugin that includes features for using geometric data (such as citizen annotations) for generating segmentation masks.
- Zooserver is a tool for managing, augmenting, and serving data on the fly to Zooniverse projects.
- Zookeeper is a python library for processing annotation data from citizen science platforms (e.g. Zooniverse) or directly from annotated images. Including tools for data extraction, clustering, masking, and visualisation.
Inspiring the next generation
The Franklin’s Science Scribbler projects also have had an impact in schools. These have been used in the classroom as part of the Zooniverse in Schools and Virus Factory in Schools programmes. From 2020 to 2023, over 50 workshops were delivered to over 500 students, by collaborators at the University of Oxford through Zooniverse in Schools. Since 2024, the Franklin’s Virus Factory in Schools programme has delivered 4 student workshops and one teacher training session. Future teacher and school activities are scheduled for the 2025/2026 school year, in collaboration with STFC, the Spires Science Curriculum Hub, and the University of Oxford. These schools engagement projects expand students’ understanding of science.
Why the Franklin?
Though citizen science is gaining traction in the public and academic communities, application of citizen science for biomedical research remains limited. The expertise developed by the Franklin staff in this field has been essential in designing successful and powerful projects in recent years, and it is this track record that continues to attract prospective collaborators.
Understanding both the biological and computational context has been essential in this work. Core expertise in the Zooniverse platform, image segmentation, machine learning, and data validation has enabled the translation of citizen effort to usable segmentation models.