Liquid Phase Electron Microscopy
Novel sample cells for electron microscopy using sheets of graphene have been developed and fabricated. These enable liquid droplets to be encapsulated for the study of biomolecules in solution at high resolution. The new cells are compatible with all standard sample holders and can be used in any transmission electron microscope. They will allow monitoring of the dynamics of biomolecules and their interactions in intact cells.

Background
To study biological molecules at the highest resolution in the transmission electron microscope it is necessary to either chemically fix or to freeze the samples. However, prevents studies of structural dynamics in biological processes. To move from static to dynamic information requires imaging in liquid environments that mimic those found in vivo. Technologies to do this will provide new insights into the dynamics of biological function.
The electron microscope operates at high vacuum and it is necessary to design sealed electron transparent cells to move this technology forward. To address this challenge, Franklin scientists looked to bring advances in material science, where Liquid-Phase Electron Microscopy (LPEM) has advanced dramatically over the past 15 years, to the life sciences. However, these techniques currently still have drawbacks including producing images with both lower contrast and lower resolution when compared with traditional electron microscopy. To unlock new biological insights, researchers must improve the design of the enclosing environment to reduce electron scattering and increase electron transparency.
Science in detail
The Franklin has developed a new generation of liquid cells for the Transmission Electron Microscopy (TEM) based on encapsulation of liquid droplets between two sheets of graphene. This technology provides an upper and lower electron transparent window with the two sheets of graphene forming a near hermetic seal around the liquid.
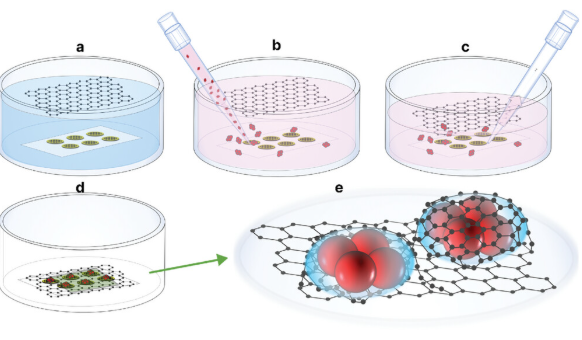
Conventional liquid cells for TEM use Microelectromechanical Systems (MEMS) fabricated silicon nitride (SiN) windows sealed with O Rings to form a liquid cell and typically have a total thickness along the electron beam direction of 50nm. The Franklin graphene cells have single carbon layer windows which reduces the electron scattering in the windows. The new cells can be fabricated in two geometries. First is a “blanket’ wherein the two sheets of graphene form a pocket encapsulating the biological structure of interest within a thin liquid film. The second is a “pocket” geometry where two graphene sheets are separated by a fabricated spacer which allows the liquid thickness relative to the diameter of the structure to be controlled.
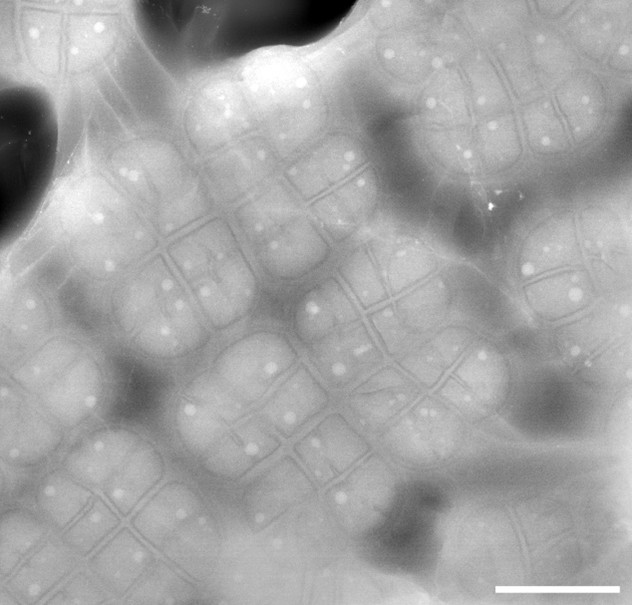
In an early proof of principle, we have used the first of these to encapsulate a prokaryotic bacterium Deinococcus radiodurans[1] and have been able to visualise the internal ultrastructure with unprecedented resolution.

Impact
The graphene liquid cell technology developed at the Franklin has already demonstrated the ability to reveal bacterial internal structures in remarkable detail at room temperature, without the need for additional sample preparation techniques that can damage or alter the structure of the specimens. This advance will enable native visualisation of organic structures at both the microscale and nanoscale. It can be generalised readily to other virus and bacterial preparations. One such capability is already being applied to visualise drug uptake and to study bacterial responses to antibiotics.
The goal of this project is ultimately to be able to image cells in liquid and to observe cellular dynamics and protein-protein interactions.
Why the Franklin?
The research required the availability of advanced EM instruments developed at the Franklin in collaboration with JEOL. Together with in-house expertise in graphene fabrication and sample preparation and biological expertise with a range of viral, bacterial and cell systems. The Institute’s interdisciplinary environment also brought together scientists from diverse fields, including materials science and biology, to tackle the challenges of liquid-phase electron imaging of biological samples and engineered liquid environments.


