
Crowd-powered research
We have developed open-source tools to support the aggregation, visualisation, and analysis of citizen science data, and shown that citizen science can produce valuable annotations for training and finetuning models for complex 3D biological imaging datasets.
Over 18,000 registered volunteers have contributed 3.49 million classifications to Franklin citizen science projects, totalling 18,000 hours of human effort since 2020. This has accelerated algorithm development, reduced individual effort through aggregation of multiple annotators, and supported public engagement and public participation in Franklin research.
These “Science Scribbler” projects have been showcased at 4 open days, 3 science festivals, and over 50 school workshops. Taking part in these events gives visitors a chance to connect with and contribute to imaging science.
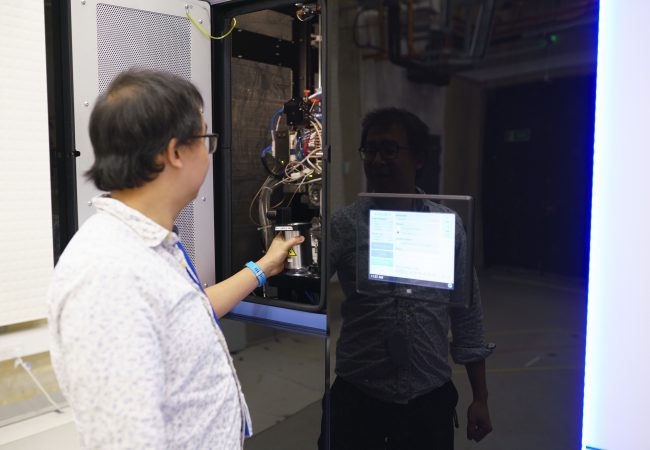

Experimental Automation
We integrate in situ structural biology, computer vision and research software engineering to develop imaging feedback–driven workflows for advanced microscopy. Our goals are to make high-resolution structural methods faster, more robust and accessible.
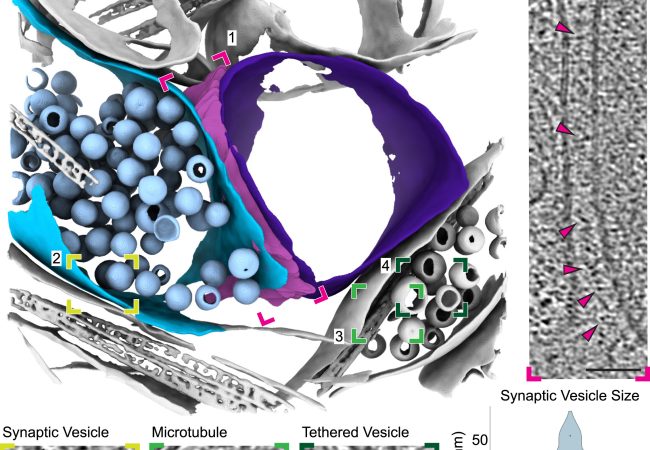
Moving from cells into tissue: a new tomography method
Researchers led by Dr Michael Grange at the Rosalind Franklin Institute have developed a robust molecular imaging workflow for brain tissue samples. Their approach utilises plasma ion beams, which increased the size of samples that could be extracted for imaging,…

Mark Basham
Mark’s primary research contributions have focused on the removal of barriers between computational methods, such as AI and data management, and scientific domains where they can be applied, alongside the open development of these techniques. He is a strong advocate…
SciCat
A community driven scientific metadata catalog designed to simplify metadata management, enabling data sharing, discovery, and collaboration.

SuRVoS2
Volume segmentation workbench offering shallow and deep approaches to segment across volumetric imaging modalities.

Red Lionfish
Richardson-Lucy deconvolution for scientists and engineers.

Ot2Rec
Automating the workflow to preprocess, reconstruct tomograms, and sub-tomogram average proteins in cryo-electron data.

AntigenApp
Web-app for managing nanobody data.

OkapiEM
A napari plugin for processing serial pFIB-SEM data.
