The Rosalind Franklin Institute scientists working as part of an international team have published the first description of an enzyme that can be used in the development and synthesis of new antiviral drugs.
The enzyme, known as ForT, plays a key role in the formation of a carbon-carbon (C-C) bond in a natural product called formycin, which has antiviral properties.
C-nucleosides like formycin are of particular interest at present following the success of the compound remdesevir as a treatment for coronavirus diseases including COVID-19, SARS and MERS.
Scientists may in future be able to re-engineer ForT’s structure to accept a greater range of substrates and produce compounds that could form the basis of new antiviral drugs.
The paper is published in the journal Chemical Communications.
Professor James Naismith, Director of the Rosalind Franklin Institute and Professor of Structural Biology at Oxford University, is one of the study’s senior authors. He said: ‘One of the key challenges in organic chemistry is making carbon-carbon bonds – it’s a difficult thing to do from a chemical perspective.
‘What’s interesting here is that some bacteria have evolved a strategy to make C-C bonds naturally, rather than the expected carbon-nitrogen (C-N) bonds, to make compounds that kill other bacteria. To help them do that, they’ve developed a new class of enzyme. Our paper is the first structural description of one of the enzymes in this class. Our finding is an important result in the field of C-nucleoside biosynthesis and an interesting insight into natural strategies for producing new molecules.’
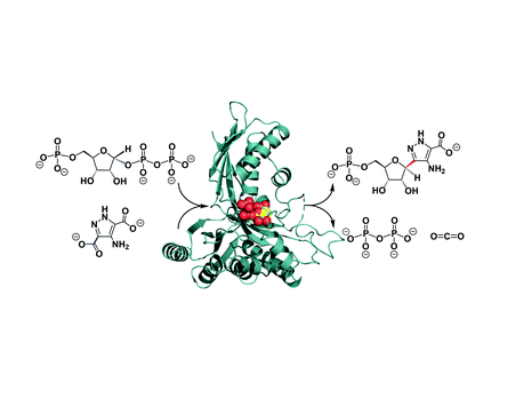
Structural biologists on the research team were able to purify the enzyme, grow crystals with the help of a substance called PRPP, and identify its active site – the area in which molecules known as substrates bind to the enzyme and undergo a chemical reaction. Data was collected using beam facilities at the Diamond Light Source synchrotron in Oxfordshire.
The team also found structural similarities with another enzyme called homoserine kinase, which catalyses a very different type chemical reaction. This unexpected result sheds light on how new enzymes might be created in evolution.
Co-senior author Professor Nigel Richards, an enzymologist based at Cardiff University’s School of Chemistry, said: ‘Until recently very little was known about how formycin is made, so our first step as enzymologists was to obtain the genetic information that would tell us how this molecule was made by certain types of bacteria. We then began collaborating with Jim on solving the crystal structures of the most interesting enzymes that are used to make this natural product.
‘Our new paper looks at the enzyme that assembles the two precursors of formycin, which is particularly interesting because the connection is a C-C rather than a C-N bond.’
Professor Richards added: ‘The chemistry catalysed by ForT gives new insights into how nature makes C-C bonds. Our findings will be of interest to the pharmaceutical industry because this enzyme provides a simplified, and therefore potentially cheaper, alternative to the chemical synthesis of C-nucleosides in the laboratory. With some re-engineering, we will try to adapt ForT, and other enzymes in this new class, to obtain new C-nucleosides that can be tested for their antiviral properties.’
This paper is part of an ongoing collaborative project funded by the BBSRC and involving groups at the Franklin and Cardiff. Their ultimate goal is to uncover the chemistry that is involved in the biosynthesis of formycin. This knowledge can then be used to develop cheaper methods of obtaining C-nucleoside antiviral drugs by chemical synthesis.
Related publication
Gao et al. - Uncovering the chemistry of C–C bond formation in C-nucleoside biosynthesis: crystal structure of a C-glycoside synthase/PRPP complex - Chemical Communications 56 (2020)