A faster route from nanobody discovery to functional testing
Researchers at the Rosalind Franklin Institute have established a faster and easier protocol to produce, test and purify large numbers of nanobodies—recombinant single-domain antibodies found in camelid species that are increasingly used in biomedical research and therapeutic development.
Published in Bio‑protocol, the method describes a streamlined, high throughput workflow for taking nanobodies identified through phage display and producing them in mammalian cells for rapid screening. By easing a common bottleneck between identifying “hits” and running functional assays, it helps researchers progress more quickly while reducing cost and material use.
Nanobodies are small antibody fragments originally derived from the immune systems of camelids such as llamas and alpacas. Unlike conventional antibodies, which are made up of multiple protein chains, nanobodies consist of a single, compact binding domain.
Their small size and stability make nanobodies versatile, and they can be combined into multimers for increased effectiveness. They are used in structural biology, diagnostics and imaging, and are increasingly being explored as the basis for new therapies. Since the COVID‑19 pandemic, nanobodies have attracted particular attention for their potential as antiviral therapeutics.
Nanobodies are typically identified by screening libraries generated either through camelid immunisation or by synthetic design. However, turning large numbers of early-stage candidates into purified, testable reagents can be slow and resource intensive, especially when projects generate hundreds of potential binders. The protocol was developed in response to a real-world bottleneck: in one project the team generated more than 400 nanobody sequences, which needed to be narrowed down to fewer than 80 candidates for deeper testing.
“We were interested in making the nanobody synthesis process as high‑throughput as possible whilst also keeping the costs low. During one project, we had so many hits after our first screen that we had to come up with a new way to process them outside our usual pipeline. Everything had to be done on a smaller scale to avoid wasting reagents and materials, as well as running multiple in parallel to manage the number of hits we were working with. Just by changing little sections of the process and adding in an extra screening step made it much more efficient and manageable.”
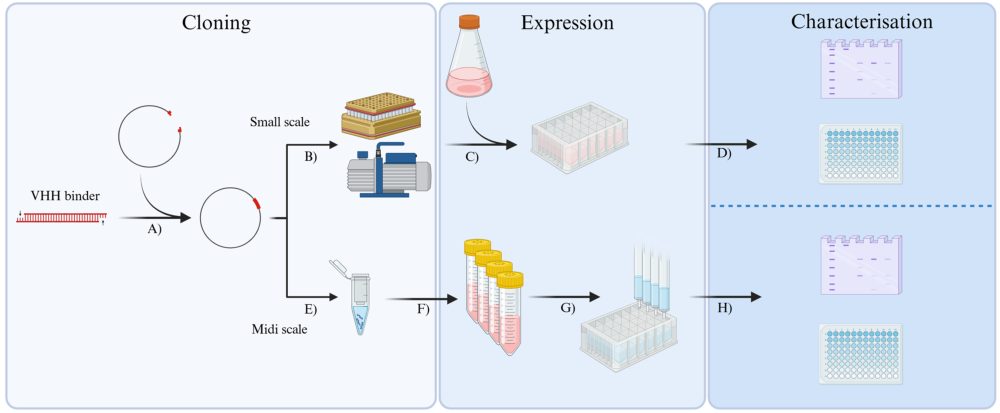
To make that possible, they adapted existing methods into a more parallel, miniaturised workflow—using expression in mammalian cells and purification steps that can be run side‑by‑side. The protocol is designed to be transferable between different nanobody formats, including Fc fusion versions often used in virus neutralisation studies, helping teams move from phage display hits to testable reagents with fewer bottlenecks.
Starting from candidate nanobodies, the workflow enables the rapid identification of those nanobodies that express well and show strong target binding. These are then purified at medium scale and taken into downstream functional studies. In order to validate the workflow, the researchers applied the method to identifying nanobodies that bind to the spike proteins of SARS‑CoV‑1 and SARS‑CoV‑2 viruses and block infection of cells.
By bringing expression in mammalian cells and purification into a single, parallel workflow, the method speeds up the pipeline between nanobody discovery and experimental application. The team hope this will support other researchers looking to move promising binders more efficiently into downstream studies.