From mice to rhinos: Combining imaging techniques to quantify 3D mammalian placentas
A cross disciplinary collaboration led by the University of Southampton with researchers at the Rosalind Franklin Institute demonstrates the power of a multi-scale, multi-modal approach to imaging and quantifying aspects of placental structure across mammals of vastly different sizes, ancestries and evolutionary strategies.
The placenta is one of the most structurally diverse organs in biology, it has evolved separately over 100 times across vertebrates, yet clinicians and biologists often rely on proxy measures such as weight, diameter or cotyledon counts to describe variation. These are easy to collect but can hide the real determinants of function- exchange surface area, exchange volume, and vascular network architecture. The team have developed new approaches to imaging and image analysis to capture these features directly, enabling like for like comparisons between species to understand the structure-function relationship in placental evolution.
The study, published in Placenta, argues that traditional, 2D or qualitative descriptions of placental form can miss key features that underpin health and disease. By combining whole organ X ray microfocus tomography (microCT), histology, and volume electron microscopy, the team delivers a holistic, quantitative “morphome” of the placenta – linking what’s seen at the organ scale to the cellular and ultrastructural levels. Researchers also can leverage the strengths of multiple techniques and minimise their weaknesses by integrating correlative imaging workflows.
The Franklin team developed and applied deep learning-based workflows for segmentation, a process that involves assigning each pixel within an image to a specific category, such as placental tissue or vasculature. This enables researchers to extract quantitative data about different elements within the image, leading to a much more robust analysis. Manually segmenting images can be extremely time-consuming, so the introduction of deep learning techniques that automate this task is an important step to make this analysis possible at scale.
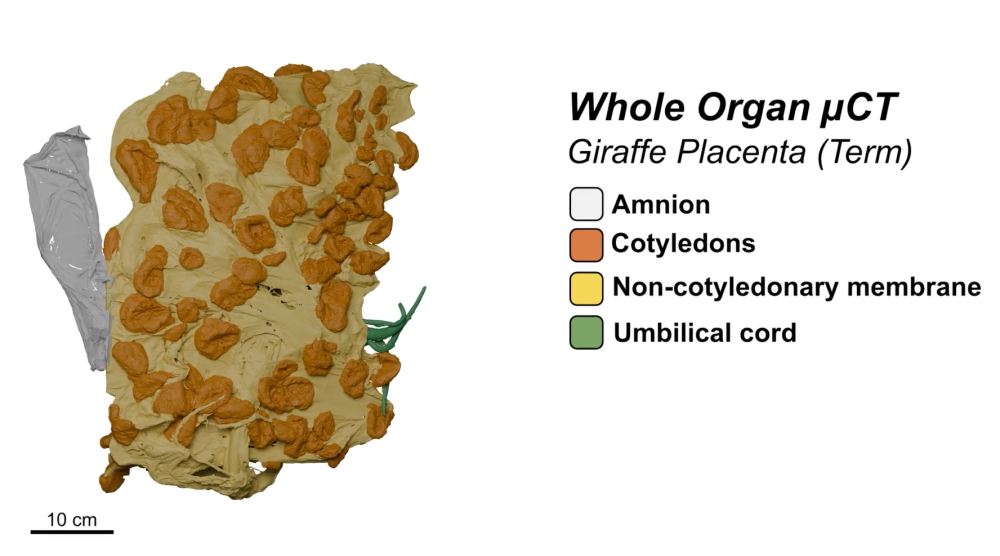

Dr Michele Darrow, Quantifying Biology across Scales Challenge lead at the Franklin, said “The placenta is an exciting and challenging area of study for the deep learning-based image segmentation workflows we have developed, which allow us to gain both quantitative measure as well as qualitative from the placental images. It has been an interesting project to learn how to adapt our workflows to the structural differences of the various species.”
Professor Rohan Lewis, Placental and Integrative Physiology Professor at University of Southampton, said “The placenta is incredibly diverse across mammals, but until recently we lacked the tools to measure that diversity properly. By combining advanced imaging with new analysis approaches, we can now map placental structure from the whole organ down to individual cells. These approaches don’t just help us understand placental evolution, they can also be applied much more widely to study complex biological structures in health and disease.”
