Life Science Challenges
Our Life Science Challenges give focus to and help drive forward our Technology Innovation Challenges. We want to create technologies which solve real world problems, so that the benefits to the research community can be maximised.
"Our primary goal is to provide technological innovation for discovery and development of new diagnostics and treatments that can improve lives worldwide."
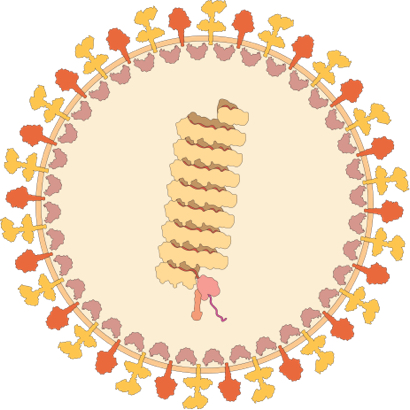

How Pathogens Interact with Human Cells
Our aim: To discover new ways of detecting, preventing and combatting human infectious diseases by discovering the mechanisms by which viruses and bacteria interact with human cells and tissues.
Emerging Interest Area: Cell-cell Interactions
The Franklin’s Emerging Interest Areas are developing areas of research led by our talented emerging leaders. These areas align with the Franklin’s mission of accelerating life science discovery and improving human health.
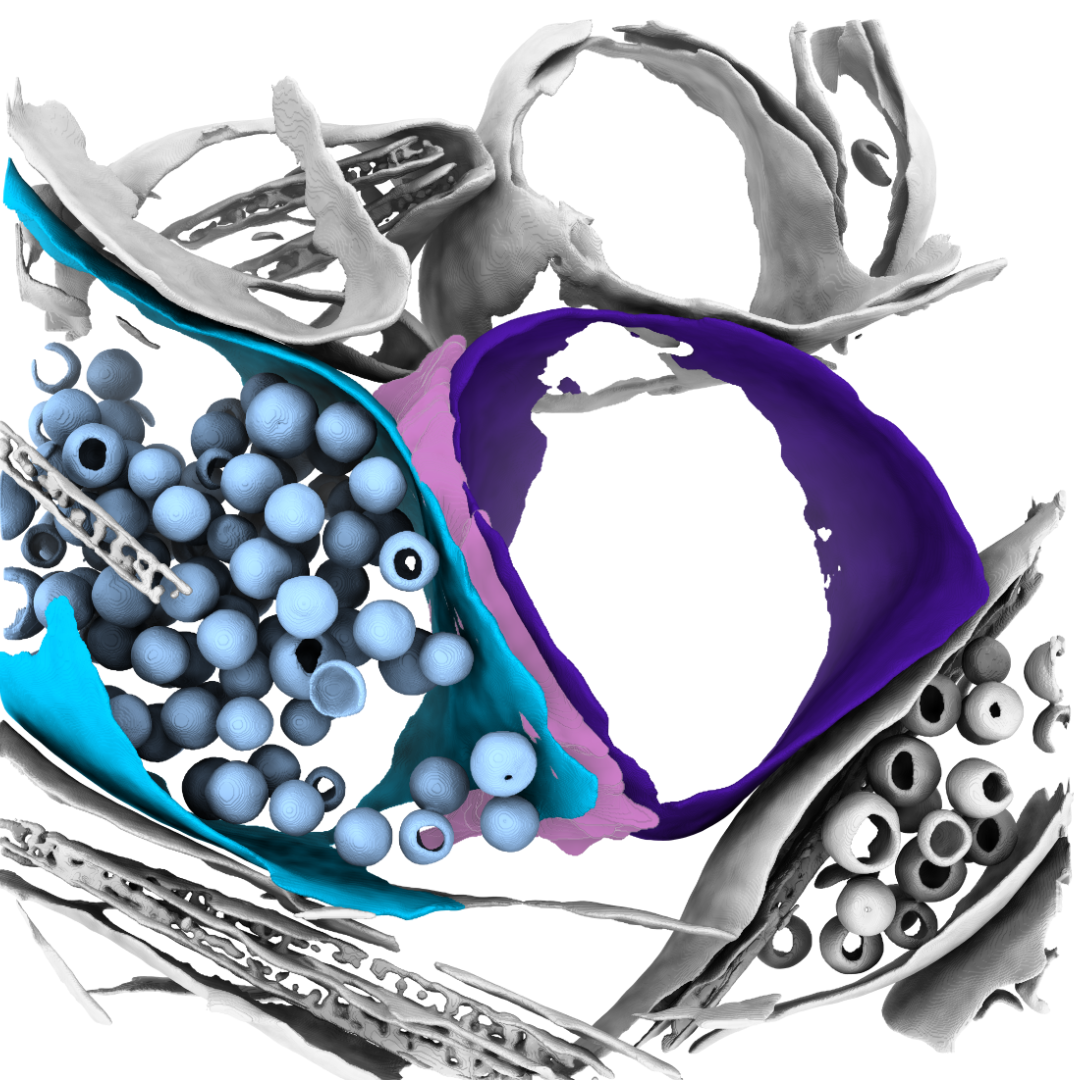

Emerging Interest Area: Structural Cell Pathology
The Franklin’s Emerging Interest Areas are developing areas of research led by our talented emerging leaders. These areas align with the Franklin’s mission of accelerating life science discovery and improving human health.

